RESEARCH
Virus-induced gene silencing (VIGS) in Cannabis sativa L.
Julia Schachtsiek, Tajammul Hussain, Khadija Azzouhri, Oliver Kayser and Felix Stehle
*Abstract
Background: The raised demand of cannabis as a medicinal plant in recent years led to an increased interest in understanding the biosynthetic routes of cannabis metabolites. Since there is no established protocol to generate stable gene knockouts in cannabis, the use of a virus-induced gene silencing (VIGS) method, resulting in a gene knockdown, to study gene functions is desirable.
Results: For this, a computational approach was employed to analyze the Cannabis sativa L. transcriptomic and genomic resources. Reporter genes expected to give rise to easily scorable phenotypes upon silencing, i.e. the phy- toene desaturase (PDS) and magnesium chelatase subunit I (ChlI), were identified in C. sativa. Subsequently, the targets of specific small interfering RNAs (siRNAs) and silencing fragments were predicted and tested in a post-transcriptional gene silencing (PTGS) approach. Here we show for the first time a gene knockdown in C. sativa using the Cotton leaf crumple virus (CLCrV) in a silencing vector system. Plants transiently transformed with the Agrobacterium tumefaciens strain AGL1, carrying the VIGS-vectors, showed the desired phenotypes, spotted bleaching of the leaves. The success- ful knockdown of the genes was additionally validated by quantitative PCR resulting in reduced expression of tran- scripts from 70 to 73% for ChlI and PDS, respectively. This is accompanied with the reduction of the chlorophyll a and carotenoid content, respectively. In summary, the data clearly demonstrate the potential for functional gene studies in cannabis using the CLCrV-based vector system.
Conclusions: The applied VIGS-method can be used for reverse genetic studies in C. sativa to identify unknown gene functions. This will gain deeper inside into unknown biosynthetic routes and will help to close the gap between avail- able genomic data and biochemical information of this important medicinal plant.
Keywords: Virus-induced gene silencing, Cannabis sativa, Cotton leaf crumple virus (CLCrV), Post-transcriptional gene silencing (PTGS)
© The Author(s) 2019. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creat iveco mmons .org/licen ses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creat iveco mmons .org/publi cdoma in/
zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
Background
Cannabis sativa L. is currently undergoing a renais- sance in the treatment of various disease symptoms [1]. Recently, in many western countries, cannabis was legalized for medicinal purposes. However, due to the long-term worldwide cultivation ban of the plant, the systematic characterization of cannabis and their metab- olites was neglected. In the last years many genomic and transcriptomic data became available [2,
3] but thefunctional characterization of gene functions lags behind.
Up to date, only a few gene functions are evaluated. One approach which could accelerate the functional charac- terization of cannabis genes is the development of a relia- ble transformation protocol, which is still challenging [4].
However, first successful hemp regeneration was recently described using novel synthetic cytokinin derivatives [5].
This would enable the use of the CRISPR–Cas9 system to produce loss-of-function mutants. One possibility to study gene functions without using stable transforma- tion protocols represent RNAi and virus-induced gene silencing (VIGS) approaches inducing transient gene knockdowns. Both have emerged as the most simple and
Open Access
*Correspondence: felix.stehle@tu-dortmund.de
Laboratory of Technical Biochemistry, Department of Biochemical and Chemical Engineering, TU Dortmund University, Dortmund, Germany
straightforward methods for reverse genetics to charac- terize functional target genes.
We utilized the Cotton leaf crumple virus (CLCrV) [6]
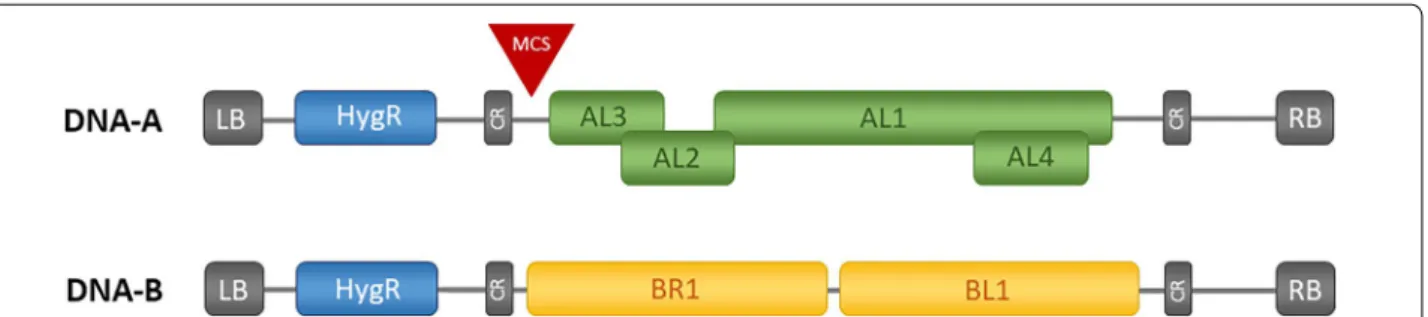
for VIGS in C. sativa. The CLCrV is a ssDNA virus that belongs to the bipartite begomoviruses consisting of two 2.5 kb circular DNA molecules, named DNA-A and DNA-B (Fig. 1) [7]. The replication is facilitated by a so- called rolling circle mechanism in the nucleus, whereas the common region (~ 200 bp) upstream of the two bidirectional-promotors act as origin for the viral DNA replication. For replication, the host machinery is nec- essary, synthesizing single- and double-stranded DNA molecules [8, 9]. The viral DNA is mobile and is present as episomes that can move in and out of the nucleus but also between different cells, tissues, and organs. This leads to systemic infection of the plant [10, 11].
VIGS makes use of the innate plant defense system against virus infections [12,
13]. Begomoviruses tran-scribe their genomes in the nuclei [14]. Subsequently, dsRNA is synthesized by the RNA-dependent RNA pol- ymerase (RDR) in the cytosol. The dsRNA then triggers post-transcriptional gene silencing (PTGS). This leads to the cleavage of the dsRNA and the generation of small interfering RNA (siRNA) molecules [15] that are rec- ognized by the RNA-induced silencing complex (RISC) [16, 17]. Subsequently, the double-stranded siRNAs are melted into single-stranded ones and those are used by the RISC to screen for complementary sequences. The resulting dsRNA molecules i.e. complexes of siRNA and virus-ssRNA will then be degraded and the replication of the virus is slowed down. This results in an amplifi- cation of the siRNA enabling a transport throughout the plant. Therefore, silencing can be observed in dis- tant cells and often in most parts of the plant [18]. VIGS exploits this RNA defense mechanism by mimicking a virus infection based on the delivery of artificial siRNA molecules to a plant cell. Thus, plant-specific mRNA molecules can be targeted resulting in a suppression of the gene expression and a reduction of the amount of the corresponding encoded proteins.
We aimed for a straightforward method to study the function of genes in C. sativa. Therefore, we established a VIGS system utilizing the Cotton leaf crumple virus (CLCrV) [6]. To visualize the knockdown, we used two maker genes resulting in a visible phenotype. First, the endogenous phytoene desaturase (PDS) gene was tar- geted. The PDS encodes an enzyme catalyzing the first step in the carotenoid biosynthetic pathway. The knock- down of the transcript will result in white leaves due to a photobleaching effect [19]. This is the result of negative feedback regulation, inhibiting the chlorophyll biosyn- thesis [20]. Second, the magnesium chelatase subunit I (ChlI) was selected. The magnesium chelatase (MgCh) is the primary enzyme for chlorophyll biosynthesis add- ing the magnesium ion into protoporphyrin IX. MgCh consists of three subunits called ChlH, ChlD and ChlI.
Among them, ChlI is of significant importance in the chlorophyll biosynthesis. ChlI subunit interacts with ChlD to activate the ChlH, which thereby regulate the whole biosynthetic pathway [21, 22]. Therefore, a loss of function in ChlI inhibits chlorophyll biosynthesis result- ing in a “yellow” phenotype.
Here, we successfully demonstrated the ability of VIGS using CLCrV. This enables to unravel gene functions, as well as to get in-depth insights to explore the distin- guished biosynthetic pathway and development in this important medicinal plant.
Methods
Identification of candidate genes and the siRNAs
For the identification of the orthologues of PDS and ChlI in C. sativa a draft genome of C. sativa was downloaded from NCBI (GCA_000230575.2; [2]. Afterwards the stan- dalone blast+ for Unix operating systems, provided by NCBI, was installed and configured (https ://www.ncbi.
nlm.nih.gov/books /NBK52 640/). A database was created
with the use of BLASTDB from in-house transcriptomic resources (not published) and the draft genome. The generated database (detailed methodology of standalone blast described by Hussain et al. [23], was used to search
Fig. 1 Schematic representation of CLCrV-vectors T-DNA structure. In between the left and right border DNA-A or DNA-B are present in addition to a hygromycin resistance cassette (HygR). DNA-A of CLCrV consists of four genes (AL1-AL4) and DNA-B consists of two genes (BR1 and BL1). The genes are flanked by the common regions (CR), consisting of the origin of replication
for homologous sequences of PDS and ChlI of C. sativa with blastn.
For the prediction of siRNAs from the identified candi- date genes the publicly available bioinformatics tool pss- RNAit (https ://plant grn.noble .org/pssRN Ait/) was used with criteria described by Xu [24]. With this information silencing gene fragments of 300 to 400 base pairs with a GC content of 30% to 70% of PDS and ChlI were defined containing the predicted siRNA target sequences. For this, fragments in the 5
′untranslated regions (UTRs), 3
′UTRs and first 100 bp of the coding sequence were excluded.
Identification of cannabis reference genes
For the identification of reference genes, suitable for quantitative real time PCR in C. sativa, the cod- ing sequences from Arabidopsis thaliana L. of typi- cally used reference genes actin (NM_001338359.1), tubulin (NM_106228), GAPDH (NM_101214.4), EF1a (NM_125432.4), ubq10 (NM_178968.5), eIFa (NM_104305.4), 18S rRNA (X16077.1), ubq5 (NM_116090.3) and yls8 (AB047811.1) were downloaded from NCBI. To find the homologous sequences in C.
sativa “blastn” 2.6.0+ was used. Sequences with a query coverage of more than 80%, an e-value of 1e10-3 and a homology greater than 70%, were chosen for a “blastx”
analysis against the NCBI non-redundant (nr) database.
Gene sequences with high homology to the respective tested gene sequences were declared as homologous sequences and can be found in Additional file 1.
Evaluation of reference genes for qPCR
The identified reference genes were tested according to their stable expression in leaves to evaluate which genes are suitable for qPCR experiments. For this purpose, qPCR was performed with cDNA of three biological rep- licates of the empty vector control plants (pCottonA) and plants infiltrated with pCotton-PDS. The analysis of the most suitable reference genes was done with the tool NormFinder [25]. Used primers for each reference gene can be found in Additional file 1: Table S1.
Plant cultivation
The cultivation of Cannabis sativa L. plants from indi- vidual seeds of the variety Finola was done under long day conditions (18 h light/6 h dark, light intensity 130 µM m
−2s
−1, white light; 4000 K) at 25 °C on hydro- culture. As fertilizer, a combination of FloraGro, Flora- Micro and Flora Bloom was used (0.03% each, General Hydroponics Europe).The temperature was changed to 22 °C during the day and 18 °C in the night after infiltra- tion with A. tumefaciens.
Verification of predicted DNA sequences of PDS and ChlI
cDNA of C. sativa var. Finola was used to verify the pre- dicted gene sequences of PDS and ChlI. The sequences were amplified with PCR and cloned into the vector pDionysos [26] by using the Gibson assembly method for better sequencing results. pDionysos was linearized with XbaI and for the amplification of PDS and ChlI the Q5
®High-Fidelity polymerase (NEB) was used. Utilized primers can be found in Additional file 1: Table S3. After Gibson cloning, constructed plasmids were sequenced to verify the predicted gene sequences.
Vector construction for agroinfiltration
For establishment of VIGS in C. sativa the Cotton leaf crumple virus (CLCrV) was applied. As a basis for the construction of the VIGS vectors the plasmids pJRT.
Agro.CLCrVA.008 (pCottonA) (Addgene plasmid # 31809) carrying DNA-A and pJRT.Agro.CLCrVB1.3 (pCottonB) (Addgene plasmid # 37974) harboring DNA-B of the virus were used. Both plasmids were a gift from Niki Robertson [6].
Construction of the VIGS-vectors was done by line- arizing the plasmid pCottonA with SpeI and cloning the selected 300 to 400 base pair fragments of PDS or ChlI into the mentioned linearized vector with the Gibson Assembly method. The resulting plasmids pCottonA- PDS and pCottonA-ChlI were sequenced to verify the correct assembly. All used primers are listed in Addi- tional file 1: Table S3.
Agroinfiltration of C. sativa
Competent A. tumefaciens LBA4404, GV3101 and AGL1
cells were transformed with the constructed plasmids
according to a published protocol [27]. For agroinfiltra-
tion the transformed A. tumefaciens cells were culti-
vated in a 5 mL-preculture in YEB medium overnight at
28 °C for strains LBA4404 and GV3101 and 26 °C for
AGL1 and 200 rpm with 50 mg L
−1kanamycin as well
as respective antibiotics for each strain (Additional
file 1: Table S2). The cultivation of the main cultures was
done according to an existing protocol [28] with minor
changes. For instance, cells from the preculture were
inoculated in 30 mL YEB medium with 50 mg L
−1kana-
mycin and respective antibiotics for each strain as well as
10 mM MES and 20 µM acetosyringone. The cells were
grown overnight at 26/28 °C and 200 rpm. For infiltra-
tion cells were harvested by centrifugation and washed
twice in infiltration buffer (10 mM MES, 10 mM MgCl
2,
200 µM acetosyringone). The cells were incubated at
room temperature for 4 h in the dark, after the OD
600nmwas adjusted to 4. After incubation, cells containing
pCottonB were mixed in a ratio of 1:1 with cells contain-
ing DNA-A with the desired gene fragments of PDS or
ChlI. The mixture was infiltrated into C. sativa var. Finola seedlings at the 4-leaf stage in the cotyledons and pri- mary leaves with a needleless syringe. After infiltration the plants were cultivated at 22 °C in the light and 18 °C in the dark.
cDNA synthesis and quantitative real‑time PCR (qPCR)
Harvested plant leaves were immediately frozen in liquid nitrogen and total RNA was extracted (NucleoSpin RNA plus Kit; Macherey Nagel). For cDNA synthesis (LunaS- cript
®RT Supermix Kit; NEB), 800 ng of DNase digested RNA were used. The cDNA was diluted to 5 ng µL
−1and qPCR was performed in five biological replicates with the Luna
®Universal qPCR Master Mix (NEB) in 20 µL reactions according to manufacturer’s instructions. The following parameters were used for amplification: 95 °C denaturation for 60 s; 40 cycles of denaturation at 95 °C for 15 s and extension at 60 °C for 30 s. Afterwards, melt- ing curves were measured. Calculation of C
tvalues was done with the StepOne V2.2.2 (Applied Biosystems) soft- ware. The data was analyzed with the use of the ∆∆C
t- method [29]. Normalization of the samples was done by using the mean of the C
tvalues of UBQ5 and eIFa. The expression level of PDS and ChlI were set to 1.0.
Proof of virus infection
To test, if the virus is present in the plants, genomic DNA was isolated five weeks after infiltration (Nucleospin Plant II, Macherey–Nagel). Detection of Virus DNA-A and DNA-B was performed in PCR experiments with the use of virus gene specific primers, respectively. As negative control genomic DNA from wild type plants was used. Amplification was done using the Red Taq DNA Polymerase Mastermix (VWR). Primers can be found in Additional file 1: Table S3.
Quantification of pigments
The extraction and quantification of photosynthetic pig- ments (chlorophyll a, chlorophyll b and carotenoids) was done according to an existing protocol [30] with minor changes. 50 mg of grounded, frozen leaf material was used for extraction of pigments with acetone. The ace- tone phase was filled up to 10 mL and 2 mL of the extract were filtered through a 0.45 µm nylon filter into a quartz cuvette. Photometric measurements were performed at 662, 645 and 470 nm.
Results
Identification of candidate genes and siRNAs prediction
For the establishment of VIGS in C. sativa, it is favora- ble to use target genes, which lead to a visible phenotype, if their transcript level is downregulated. Suitable genes, already used in many other VIGS approaches, represent
the genes encoding for the phytoene desaturase (PDS) and magnesium chelatase subunit I (ChlI). Both genes involved in the photosynthesis/carotenoid biosynthesis and silencing of these genes lead to white/yellow leaves phenotype. Since these genes were not identified from C. sativa yet, homologous sequences need to be iden- tified from the cannabis draft genome and transcrip- tomic resources. For this purpose, reference sequences from AtPDS and AtChlI of Arabidopsis thaliana L. were used. To search for homologies, a command-line stan- dalone BLAST+ for Linux (version 2.6.0+) was used.
Afterwards, a database for the cannabis draft genome and the transcriptomic data was created by utilizing the “BLASTDB” algorithm, and with the use of “blastn”, respective homologous sequences were found (Fig.
2a).Sequences with at least 60% identity and a stringent e-value of 1e−05 were considered for further analysis.
For verification of the candidate sequences, they were compared with the use of BLAST+ against NCBI non- redundant protein (nr), nucleotide (nt) and Swiss-Prot (UniProtKB) databases with the blastx algorithm. Only sequences with significant similarity of over 60% with the reference sequences of ChlI and PDS from other plant species were considered as potential homologs in C.
sativa. Both identified sequences (Cs-PDS and Cs-ChlI) share over 80% homology at nucleotide and protein level compared to the reference sequences from Arabidopsis.
Validation of the computationally predicted sequences was carried out by PCR amplification. Then the amplified coding sequences were cloned into a vector using Gib- son assembly. Afterwards they were verified by Sanger sequencing. The results showed a significant similarity of 99.9% in comparison to the predicted sequences. Both sequences were submitted to NCBI (GenBank accession numbers: CsPDS, MN395698; CsChlI, MN395699) and can be found in Additional file 1.
To achieve efficient gene silencing it is necessary to select gene fragments that are able to form efficient siR- NAs thereby possessing minimal off-target silencing effects. For this purpose, the identified sequences were analyzed by using the bioinformatics tool “pssRNAit”
(https ://plant grn.noble .org/pssRN Ait/). The tool pre- dicted 21 siRNAs and 347 off-targets for Cs-PDS and 16 siRNAs and 259 off-targets for Cs-ChlI. To filter out off-target sites, which might have other binding sites other than Cs-PDS and Cs-ChlI, all found siRNAs were scanned against the cannabis database, described above.
Finally, one siRNA was selected for each gene to design
the silencing gene fragments (SGF). Fragments were con-
sidered, having a length of 150–400 base pairs and a GC
content between 30 and 70%. The designed SGF for PDS
had a length of 392 base pairs and the SGF chosen for
ChlI had a fragment size of 300 base pairs.
Cotton leaf crumple virus is suitable to induce PTGS in cannabis
We chose the Cotton leaf crumple virus (CLCrV), a cir- cular ssDNA virus for VIGS experiments in C. sativa.
At first, the selected PDS fragment was cloned into pCottonA for the generation of the plasmid pCotton- PDS, which was transformed into the three A. tume- faciens strains for evaluation of the best-performing strain. The DNA-B virus component (Fig.
1), locatedon the plasmid pCottonB and the empty DNA-A vector (pCottonA), were transformed into the same Agrobac- terium strains. C. sativa plants were infiltrated either with pCotton-PDS or pCottonA as an empty-vector control in combination with pCottonB in a ratio of 1:1 (Fig.
2b). Plants were infiltrated with Agrobacteria atthe 4-leaf stage with a needleless syringe, 7 to 10 days after germination into cotyledons and first leaves.
In the beginning, 16 plants were infiltrated with each Agrobacterium strain. The infiltrated plants were grown in climate chambers and four weeks after infiltration leaves of two plants infiltrated with the strain AGL1 showed the desired phenotype of bleached leaves, indi- cated by small white spots (Fig.
3a). Control plantsshowed no phenotype. Notably, no phenotype could be observed for plants infiltrated with strains GV3101 and LBA4404 up to this point. Thus, it was suggested that using the strain AGL1 for infiltration of the plants is more effective with regard to gene silencing and further infiltrations were only carried out with the Agrobacte- rium strain AGL1. In total, 38 plants were infiltrated with the strain AGL1 carrying pCotton-PDS, whereof 13 plants died within 3 days. From the remaining 25 plants, five plants showed the desired phenotype on
newly developed leaves, leading to an overall efficiency of 20% (Fig. 3b).
Validation of the knockdown of PDS with qPCR
The white-dotted leaves of the pCotton-PDS-infiltrated plants indicate a downregulation of the transcript level of the PDS gene. However, no completely white leaves were observed. To investigate to which extend the transcript of PDS was downregulated in the described plants, quan- tification of the transcript level of PDS was done with quantitative real-time PCR (qPCR). For this purpose, it was necessary to find a suitable combination of refer- ence genes. Nine reference genes could be identified from the cannabis transcriptomic data, which should be tested for stable expression either in control plants and plants infiltrated with pCotton-PDS, including the commonly used candidates actin2, tubulin1, 18S rRNA, glyceralde- hyde 3-phsophate dehydrogenase (GAPDH), elongation factor 1α (EF1α), eukaryotic initiation factor a (eIFa), ubiquitin 5 and ubiquitin 10 (ubq5, ubq10). Additionally, the gene yellow leaf specific protein 8 (YLS8) was consid- ered, which was found to be a suitable reference gene in hop (Humulus lupulus L.) [31], a plant belonging to the Cannabaceae family as well. After isolation of RNA from leaves of three biological replicates of the empty vector control plants and plants infiltrated with pCotton-PDS, cDNA was synthesized and used for qPCR experiments.
Each gene was analyzed in three technical replicates. To
find the most stable reference gene, the tool NormFinder
was used [25], which allows a prediction of the best ref-
erence gene or best combination of reference genes,
by analyzing the C
Tvalues of the samples. Most stable
expression, i.e. lowest stability values were obtained for
Fig. 2 Workflow of a VIGS experiment a after identification of gene sequences and validation by sequencing, suitable siRNAs and off-targets are predicted. Based on these predictions silencing gene fragments were defined. b For in vivo validation silencing fragments are cloned in the VIGS-vectors and transformed in Agrobacteria for infiltration experimentsUBQ5 and eIFa (Additional file 1: Table S4). Thus, they were used as reference genes for normalization in all fol- lowing qPCR experiments.
In comparison to the control plants, infiltrated with the empty vector pCottonA, the transcript level of PDS was downregulated by 73% in plants (Fig. 3c), which were infiltrated with pCotton-PDS and showing a visible phe- notype, as described above (Fig. 3a). This indicates, that the observed phenotype is linked to the downregulated transcript level and gene silencing of the PDS gene was possible with the predicted silencing fragment and the related predicted siRNA. Furthermore, this was con- firmed by quantification of the pigments (chlorophyll a, chlorophyll b and carotenoids). Leaves with a visible phenotype showed a decrease of chlorophyll a and carot- enoids by 27% and 29%, respectively. However, the chlo- rophyll b content was not affected (Fig. 3d).
ChlI as a second marker for VIGS in cannabis
In a second approach, the developed system should be transferred to a second marker gene, showing a vis- ible phenotype as well, the ChlI gene. Since this gene is related to photosynthesis, downregulation would lead to a bleaching phenotype as well. For cotton it was observed, that targeting ChlI for gene silencing had a
greater visible bleaching effect than PDS-silencing [9].
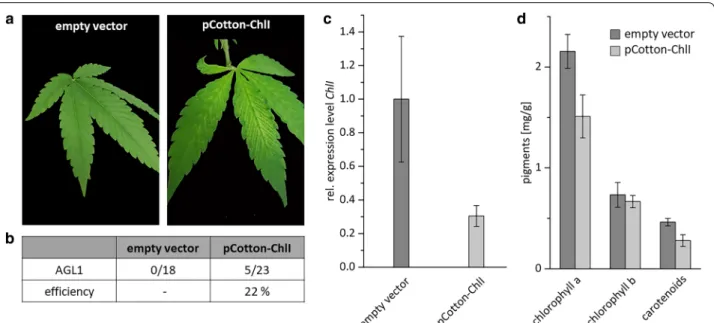
To test this, the chosen silencing fragment was cloned into pCottonA and transformed only into A. tumefa- ciens AGL1 cells. As described above, cells were culti- vated, harvested and the optical density was diluted to 4.0 in infiltration buffer. In total 40 plants were infil- trated with pCotton-ChlI, whereof 23 survived. As empty vector control, 8 plants were infiltrated with pCottonA. Whereas control plants showed no phe- notype at all, five plants infiltrated with the silencing construct (pCotton-ChlI) showed the desired pheno- type after approximately four weeks, representing an efficiency of 22%. As already observed for PDS silenced plants, the newly developed leaves are spotted with white dots all over the leaves. No complete bleaching was observed as well (Fig. 4a, b).
Transcript analysis of five biological replicates indi- cated a downregulation of 70% relative to the control plants infiltrated with the empty vector pCottonA (Fig.
4c). Similar to the PDS-silencing approach, thepigment content was analyzed to confirm the given results. Compared to the empty vector control plants chlorophyll a was decreased by 30% and the carotenoid content by 40% in ChlI-silenced plants.The chlorophyll b level was not affected (Fig. 4d).
Fig. 3 Silencing of PDS induced with the Cotton leaf crumple virus a Plants were infiltrated with the A. tumefaciens strain AGL1 carrying pCotton-PDS or an empty-vector control. Phenotypes could be identified approximately four weeks after infiltration with pCotton-PDS (right), whereas control plants showed no phenotype (left). b Overview on the efficiency of VIGS with different A. tumefaciens strains and number of infiltrated plants.
c Quantification of the expression level of PDS in plants showing a phenotype and empty- vector control plants were analyzed by qPCR and normalized relative to the expression levels of eIFa and UBQ5. Error bars indicate standard error of five biological replicates. d Quantification of pigment content (chlorophyll a, chlorophyll b and carotenoids) of leaves from empty-vector control plants and PDS-silenced plants. Error bars indicate standard error of 3 representative replicates
Virus is present in infiltrated plants
Finally, the presence of both virus ssDNAs (DNA-A and DNA-B) was verified in agroinfiltrated plants. For this purpose primers specific for either DNA-A (AL1 gene) or DNA-B (BL1 gene) (Fig.
1) were designed amplify-ing short fragments. eIFa was used as internal control to
ensure similar amounts of genomic DNA were used for amplification (Fig.
5). DNA-A and DNA-B fragmentswere present in plants transformed with empty vectors as well as in PDS and ChlI silenced plants, indicating both ssDNAs in all analyzed leaves.
Discussion and conclusions
At the moment no reliable protocols are available, which enable regeneration and stable transformation of canna- bis plants [4]. Nevertheless, having the ability to analyze individual gene functions of cannabis would be a great step to characterize the plant further. For this purpose, the method of virus-induced gene silencing was evalu- ated. Here we show for the first time that VIGS, can be applied to C. sativa, using the Cotton leaf crumple virus, for the downregulation of genes, by applying the virus to the plant cells transiently via agroinfiltration.
In first infiltration experiments with the Cotton leaf crumple virus and the three different A. tumefaciens strains LBA4404, GV3101 and AGL1, it could be shown, that only seedlings transformed with AGL1 showed the expected phenotype. This leads to the hypothesis that Agroinfiltration in C. sativa is only successful when using a hypervirulent A. tumefaciens strain, like AGL1 [32].
However, the observed phenotype of plants infiltrated with pCotton-PDS was not as strong as known from tobacco rattle virus (TRV)-infected plants. Only white
Fig. 4 Silencing of ChlI induced with the Cotton leaf crumple virus a Plants were infiltrated with the A. tumefaciens strain AGL1 carrying pCotton-ChlI or an empty-vector control. Phenotypes were visible approximately four weeks after infiltration with pCotton-ChlI (right). No phenotype was visible in control plants (left). b Overview on the efficiency of VIGS if plants were infiltrated with the A. tumefaciens strain AGL1. c Expression level of ChlI in plants showing a phenotype and empty-vector control plants were quantified with qPCR and normalized relative to the expression levels of eIFa and UBQ5. Error bars indicate standard error of five biological replicates. d Quantification of pigment content (chlorophyll a, chlorophyll b and carotenoids) of leaves from empty-vector control plants and ChlI-silenced plants. Error bars indicate standard error of 3 representative replicatesFig. 5 Detection of DNA-A and DNA-B. Genomic DNA was extracted either from PDS- and ChlI-silenced plants five week after inoculation or from control plants. As internal standard eIFa was used to ensure similar amounts of genomic DNA were used for amplification
and yellow spotted leaves instead of a complete bleach- ing of the green color was observed. Nearly the same phenotypes, i.e. white and yellow dots all over the sur- face of the leaves, were observed by applying this virus- system to the native host for gene knockdowns [6,
9].Furthermore, the visible phenotype reflects the quanti- fied pigment content, which was up to 40% lower com- pared to the empty-vector control plants.
To measure the expression levels of the targeted genes it was essential to set up a protocol for quanti- tative real-time PCR. For reliable results a combina- tion of reference genes is required that shows the most stable expression. Nine potential reference genes were identified and analyzed with the NormFinder tool [25]
resulting in UBQ5 and eIFa as the best combination of reference genes for normalization. Since the coding sequences of potential reference genes are available by now, they can be used for designing future qPCR exper- iments e.g. for other tissues than leaves or different cannabis varieties, respectively.
According to the observed phenotype, it was not sur- prising that the silencing effect of 73% and 70% in our study, respectively, was not as high as in experiments utilizing the TRV. Interestingly, the comparison of our results with the silencing effects of other studies shows that the determined transcript levels are within the same range. In N. benthamiana and tomato gene silencing of minimum 78% was reached [8, 33]. With the use of the Foxtail Mosaic Virus in barley, transcription levels were reduced by minimum 75% [34].
In total, the silencing-efficiency of PDS and ChlI with 20% and 22%, respectively, is rather low. In cotton the silencing-efficiencies were 81% and 65% for agroinfiltra- tion or particle bombardement, respectively [6]. How- ever, in N. benthamiana plants silencing-efficiencies up to 98% could be reached by using TRV [35]. This dem- onstrates that CLCrV exhibits an overall lower silencing- efficiency compared to TRV. By using a non-native host, like cannabis, reduced efficiencies can be assumed. Nev- ertheless, so far no suitable cannabis-specific virus is known, which is able to show virus-induced symptoms and exhibit the capability to move between plant cells [36].
For tobacco and flax it could already be shown, that the removal of the apical shoot meristem, right after Agro- infiltration, led to a higher silencing efficiency, due to a more effective spread of the virus within the plant [37,
38]. However, it could be shown, that the virus is ableto spread into new developing leaves, since fragments of both ssDNAs could be amplified. Since control plants showed no bleaching effect it can be concluded that the inserted fragments of PDS and ChlI were responsible for efficient gene silencing.
This shows, that despite the weak phenotype, the developed VIGS-method in C. sativa can be used to perform reverse genetic approaches to identify unknown gene functions. This will help to elucidate unknown biosynthetic routes and will contribute to a deeper knowledge of medical cannabis. Moreover, the established method is also suitable for the evalua- tion of endogenous miRNAs and their corresponding targets [39, 40]. This opens the possibility to elucidate gene functions and to decipher the regulatory network of altering transcription levels for a better understand- ing of the regulation of biosynthetic routes and plant development.
Supplementary information
Supplementary information accompanies this paper at https ://doi.
org/10.1186/s1300 7-019-0542-5.
Additional file 1. Table S1. Primers used for qPCR experiments. Table S2.
Antibiotic concentrations used for cultivation of Agrobacterium tume- faciens strains. Table S3. Primers used in the study for construction of the vectors. Table S4. Results from the NormFinder analysis of reference genes suitable for qPCR.
Acknowledgements
We acknowledge financial support by Deutsche Forschungsgemeinschaft (DFG) and Technische Universität Dortmund/TU Dortmund Technical Univer- sity (Germany) within the funding program Open Access Publishing. Studies were performed with the permission of No. 458 64 16 issued by the Federal Institute for Drugs and Medical Devices (BfArM), Germany.
Authors’ contributions
JS and FS planned the experimental studies. TH analyzed the genomic data and identified the used cannabis genes and siRNAs/silencing fragments. JS cloned the constructs. JS and KA performed VIGS and carried out qPCR experi- ments. OK is the principal investigator, FS coordinated and supervised the study. JS, TH and FS wrote the manuscript. All authors read and approved the final manuscript.
Funding
Open Access Publishing was financially support by Deutsche Forschun- gsgemeinschaft (DFG) and Technische Universität Dortmund/TU Dortmund Technical University (Germany).
Availability of data and materials
The material used during the current study are available from the correspond- ing author on reasonable request.
Ethics approval and consent to participate
Local, National and International guidelines were followed in this study with virus-induced gene silencing in plants.
Consent for publication Not applicable.
Competing interests
The authors declare that they have no competing interests.
Received: 27 September 2019 Accepted: 5 December 2019
References
1. Pisanti S, Bifulco M. Medical Cannabis : A plurimillennial history of an evergreen. J Cell Physiol. 2018. https ://doi.org/10.1002/jcp.27725 . 2. van Bakel H, Stout JM, Cote AG, Tallon CM, Sharpe AG, Hughes TR, et al.
The draft genome and transcriptome of Cannabissativa. Genome Biol.
2011;12:R102. https ://doi.org/10.1186/gb-2011-12-10-r102.
3. Laverty KU, Stout JM, Sullivan MJ, Shah H, Gill N, Deikus G, et al. A physical and genetic map of Cannabissativa identifies extensive rearrangement at the THC/CBD acid synthase loci. Genome Res. 2019;29:146–56.
4. Schachtsiek J, Warzecha H, Kayser O, Stehle F. Current perspectives on biotechnological cannabinoid production in plants. Planta Med.
2018;84:214–20.
5. Smýkalová I, Vrbová M, Cvečková M, Plačková L, Žukauskaitė A, Zatloukal M, et al. The effects of novel synthetic cytokinin derivatives and endog- enous cytokinins on the in vitro growth responses of hemp (Cannabis sativa L.) explants. Plant Cell Tissue Organ Cult. 2019;139:381–94.
6. Tuttle JR, Haigler CH, Robertson D. Method: low-cost delivery of the cotton leaf crumple virus-induced gene silencing system. Plant Methods.
2012;8:1–8.
7. Bisaro DM. Silencing suppression by geminivirus proteins. Virology.
2006;344:158–68.
8. Liu YL, Schiff M, Marathe R, Dinesh-Kumar SP. Tobacco Rar1, EDS1 and NPR1/NIM1 like genes are required for N-mediated resistance to tobacco mosaic virus. Plant J. 2002;30:415–29.
9. Tuttle JR, Idris AM, Brown JK, Haigler CH, Robertson D. Geminivirus- mediated gene silencing from cotton leaf crumple virus is enhanced by low temperature in cotton. Plant Physiol. 2008;148:41–50. https ://doi.
org/10.1104/pp.108.12386 9.
10. Noueiry AO, Lucas WJ, Gilbertson RL. Two proteins of a plant DNA virus coordinate nuclear and plasmodesmal transport. Cell. 1994;76:925–32.
11. Sanderfoot AA, Ingham DJ, Lazarowitz SC. A viral movement protein as a nuclear shuttle. Plant Physiol. 1992;110(1):23–33.
12. Baulcombe DC. Viruses and gene silencing in plants. In: Calishe CH, Horzinek MC, editors. 100 years of virology. Archives of Virology. Supple- menta. Vienna: Springer; 1999. p. 189–201.
13. Ratcliff FG, MacFarlane SA, Baulcombe DC. Gene silencing without DNA: RNA-mediated cross-protection between viruses. Plant Cell.
1999;11:1207–15.
14. Hanley-Bowdoin L, Settlage SB, Orozco BM, Nagar S, Robertson D.
Geminiviruses: models for plant DNA replication, transcription, and cell cycle regulation. CRC Crit Rev Plant Sci. 1999;18:71–106. https ://doi.
org/10.1080/07352 68999 13091 62.
15. Zamore PD. RNA interference: listening to the sound of silence. Nat Struct Biol. 2001;8:746–50.
16. Ding SW, Voinnet O. Antiviral immunity directed by small RNAs. Cell.
2007;130:413–26.
17. Waterhouse PM, Fusaro AF. Viruses face a double defense by plant small RNAs. Science (80). 2006;313:54–55.
18. Kalantidis K, Schuhmacher HT, Alexiadis T, Helm JM. RNA silencing move- ment in plants. Biol Cell. 2008;100:13–26.
19. Kumagai M, Donson J, Harvey D, Hainley K, Grill LK. Cytoplasmic inhibi- tion of carotenoid biosynthesis with virus-derived RNA. Proc Natl Acad Sci. 1995;92:1679–83.
20. Qin G, Gu H, Ma L, Peng Y, Deng XW, Chen Z, et al. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res.
2007;17:471–82.
21. Tanaka R, Tanaka A. Tetrapyrrole biosynthesis in higher plants. Annu Rev Plant Biol. 2007;58:321–46. https ://doi.org/10.1146/annur ev.arpla nt.57.03290 5.10544 8.
22. Willows RD. Biosynthesis of chlorophylls from protoporphyrin IX. Nat Prod Rep. 2003;20:327–41.
23. Hussain T, Plunkett B, Ejaz M, Espley RV, Kayser O. Identification of putative precursor genes for the biosynthesis of cannabinoid-like compound
in Radula marginata. Front Plant Sci. 2018. https ://doi.org/10.3389/
fpls.2018.00537 .
24. Xu P, Zhang Y, Kang L, Roossinck MJ, Mysore KS. Computational estima- tion and experimental verification of off-target silencing during posttran- scriptional gene silencing in plants. Plant Physiol. 2006;142:429–40. https ://doi.org/10.1104/pp.106.08329 5.
25. Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantita- tive reverse transcription-PCR data: a model-based variance estima- tion approach to identify genes suited for normalization, applied to bladder and colon cancer data sets normalization of real-time. Cancer.
2004;64:5245–50.
26. Stehle F, Stubbs MT, Strack D, Milkowski C. Heterologous expression of a serine carboxypeptidase-like acyltransferase and characterization of the kinetic mechanism. FEBS J. 2008;275:775–87.
27. Höfgen R, Willmitzer L. Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res. 1988;16:9877.
28. Burch-Smith TM, Schiff M, Liu Y, Dinesh-Kumar SP. Efficient virus-induced gene silencing in Arabidopsis. Plant Physiol. 2006;142:21–7. https ://doi.
org/10.1104/pp.106.08462 4.
29. Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations.
Biotechnol Lett. 2004;26:509–15. https ://doi.org/10.1023/B:BILE.00000 19559 .84305 .47.
30. Ullrich SF, Rothauer A, Hagels H, Kayser O. Influence of light, temperature, and macronutrients on growth and scopolamine biosynthesis in Duboisia species. Planta Med. 2017;83:937–45.
31. Štajner N, Cregeen S, Javornik B. Evaluation of reference genes for RT- qPCR Expression studies in Hop (Humulus lupulus L.) during infection with vascular pathogen Verticillium albo-atrum. PLoS ONE ONE. 2013;8:e68228 32. Lazo GR, Stein PA, Ludwig RA. A DNA transformation–competent Arabi-
dopsis genomic library in Agrobacterium. Bio/Technology. 1991;9:963–7.
33. Liu Y, Schiff M, Dinesh-Kumar SP. Virus-induced gene silencing in tomato.
Plant J. 2002;31:777–86. https ://doi.org/10.1046/j.1365-313X.2002.01394 .x.
34. Liu N, Xie K, Jia Q, Zhao J, Chen T, Li H, et al. Foxtail Mosaic Virus-induced gene silencing in monocot plants. Plant Physiol. 2016;171:1801–7. https ://
doi.org/10.1104/pp.16.00010 .
35. Tian J, Pei H, Zhang S, Chen J, Chen W, Yang R, et al. TRV-GFP: A modified Tobacco rattle virus vector for efficient and visualizable analysis of gene function. J Exp Bot. 2014;65:311–22.
36. Righetti L, Paris R, Ratti C, Calassanzio M, Onofri C, Calzolari D, et al. Not the one, but the only one: about Cannabis cryptic virus in plants showing
‘hemp streak’ disease symptoms. Eur J Plant Pathol. 2018;150:575–88.
37. Wijdeveld MMG, Goldbach RW, Meurs C, van Loon LC. Accumulation of viral 126 kDa protein and symptom expression in tobacco systemically infected with different strains of tobacco mosaic virus. Physiol Mol Plant Pathol. 1992;41:437–51. https ://doi.org/10.1016/0885-5765(92)90055 -Z.
38. Chantreau M, Chabbert B, Billiard S, Hawkins S, Neutelings G. Functional analyses of cellulose synthase genes in flax (Linum usitatissimum) by virus-induced gene silencing. Plant Biotechnol J. 2015;13:1312–24. https ://doi.org/10.1111/pbi.12350 .
39. Tiwari M, Sharma D, Trivedi PK. Artificial microRNA mediated gene silenc- ing in plants: Progress and perspectives. Plant Mol Biol. 2014;86:1–18.
40. Yan F, Guo W, Wu G, Lu Y, Peng J, Zheng H, et al. A virus-based miRNA suppression (VbMS) system for miRNA loss-of-function analysis in plants.
Biotechnol J. 2014;9:702–8.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in pub- lished maps and institutional affiliations.